Survival of Generic Escherichia coli and Salmonella in Oregon's Agricultural Soils
Abstract
Establishment of foodborne pathogens in agricultural production areas (orchards, fields) has been linked to produce-related outbreaks. The source of contaminants is often undetermined; however, agricultural water could serve as a conduit for the initial and continual contamination of produce. Of particular concern, is the potential persistence of foodborne pathogens in agricultural soils used for crops that come in contact with the ground (i.e., root crops, tree nuts). Previous research has been conducted on the survival and persistence in various soil types; however, definitive and harmonized conclusions of critical soil characteristics have not been established, making it difficult to apply these findings to other soil types. The objective of this study was to compare the relative survival and persistence of generic E. coli and Salmonella in a variety of Oregon's agricultural soils when irrigated with contaminated water. Adkins, Cullius, Latourell, Madras, Quatama, and Willamette soils were collected from state experiment stations and irrigated with inoculated irrigation water to achieve high (4 log CFU/g) and low (2 log CFU/g) contamination levels of generic E. coli and Salmonella. Soils were held under ambient conditions in a temperature-controlled greenhouse (10-24 ℃) and analyzed for surviving populations for up to 86 days post-inoculation. Soils varied significantly in the rate of reduction and persistence of bacterial contaminants, with Quatama being the most inhibitory and Cullius being the least inhibitory. These findings suggest that soils high in clay, particularly smectite, have the highest potential for supporting the survival and long-term persistence of generic E. coli and Salmonella.
Keywords
Generic E. Coli, Persistence, Salmonella, Soils, Clay content
Introduction
Approximately 50% of food borne illness outbreaks in the U.S. are linked to produce [1-3]. In an attempt to reduce the illness burden associated with produce, the U.S. Food and Drug Administration (FDA) is implementing rules meant to mitigate contamination along the production and supply chain [4,5]. Within the "Produce Safety Rule", agricultural water quality, specifically microbiological quality, has been highlighted as an area of on-farm food safety that will require increased monitoring and potential mitigation. The highest risk associated with poor quality water has been attributed to applications in which water comes in direct contact with the harvestable portion of the crop; therefore, this has been the focus during the continued evolution of water quality regulations [4,6]. While contact with water represents the most immediate concern for produce contamination, there is also the potential for poor quality water to serve as a conduit for the long-term establishment of foodborne pathogens in agricultural fields and orchards. This contamination is of particular concern for crops that come into direct contact with soil prior to harvest (i.e., root crops, tree nuts). The 5-year persistence of an outbreak-associated Salmonella Enteritidis PT30 in an almond orchard has demonstrated this as a serious risk [7,8].
As fields and orchards are irrigated with contaminated water, potentially harmful microorganisms are distributed into the soil. The persistence of relevant indicators and pathogens is dependent on many factors, including specific hydrological aspects as well as agronomic practices; however, of particular interest is the relative ability of contaminants to survive in various agricultural soil types. Previous studies indicated that foodborne pathogens, such as E. coli O157:H7 and Salmonella, survive and persist with varying ability in different soil types [9-16]. Differences in study design and storage conditions have led to various conclusions about soil characteristics that significantly contribute or predict pathogen survival and persistence.
Soils vary greatly in their physical and chemical characteristics even within fields that appear uniform; this variation influences the survival and persistence of microorganisms in soil. Oregon grows over 250 crops on a diversity of soils representing over 300 unique soil series of prime farmland [17]. Previous work demonstrated similar decay rates, but significantly different persistence of generic E. coli and Salmonella in two dissimilar agricultural soils soured from onion fields in Oregon: Semiahmoo muck (irrigation not required-Typic Haplosaprists) and Owyhee silt loam (irrigation required - Xeric Haplocalcids) [18]. The complexity and variability of soil systems as well as the lack of harmonized conclusions from these studies make it challenging to predict survival and persistence of contaminating bacteria in soil types or under moisture and temperature conditions differing from those used in the individual studies. To assist the regional agriculture industry in understanding their potential food safety risks and guide further research on mitigation strategies, it would be helpful to demonstrate the survival and persistence of indicators and pathogens in relevant food production soils.
The primary objective of this study was to compare the relative survival and persistence of generic E. coli and Salmonella in a variety of Oregon's agricultural soils when irrigated with contaminated water. Findings from these studies could guide mitigation strategies for fields that have been contaminated by water applications. Six soils were collected from dominant and diverse growing regions in Oregon to determine the relative risk of long-term contamination. The Alfisols soil order is represented by two fine-loamy soil series, Quatama and Latourell, which are in the Aquultic and Ultic Haploxeralfs subgroups, respectively. The Aridisols order is represented by the coarse-loamy Adkins soil series, which is a Xeric Haplocalcids. The remaining three soils (Willamette, Madras, and Cullius) are members of the Mollisols. Willamette is a fine-silty Pachic Ultic Argixerolls, Madras is a fine-loamy Aridic Argixerolls, and Cullius is a clayey Aridic Lithic Argixerolls [19,20]. Additional details on these soil series have been compiled in (Table 1).
Materials and Methods
Greenhouses
Two Oregon State University (OSU) greenhouses (West 6-5 - 700 sq. ft; West 7-6 - 340 sq. ft.) were used for these survival studies. Both greenhouses were constructed of solid concrete floors with steel mesh grid tables (5' × 15'). Greenhouse temperatures were maintained by thermostatic control (High: 24 ℃; Low: 10 ℃) and temperature values throughout the study were recorded using a weather station (EasyweatherProweather station, Tycon Power Systems, Bluffdale, UT). To assist with the maintenance of temperature, greenhouse windows were covered with polyethylene fabric shade cloth throughout the duration of this study. All greenhouse operations were managed by OSU greenhouse personnel.
Soils
Six soil types (225 kg each) were collected from Oregon State University Experiment Stations in diverse growing regions and transported to the Oregon State University greenhouses (Table 1). Soil samples (500 g) were submitted to Edge Analytical Laboratories (Corvallis, OR) for pH and mineral composition analysis. Soils were sifted through wire mesh grid boxes (1.27 cm grid) to reduce clumping and distributed (400g) into plastic sample cups (500 ml; Party City Corporation, Rockaway, NJ). Sample cups (n = 100) were distributed into large grow trays (4' × 4'; Botanicare, Chandler, AZ) to contain any contaminated runoff. A total of 24 trays were used and each tray was treated as a block for a given treatment (inoculation level and soil type) with the treatments randomized across both greenhouses. Polyvinyl chloride pipe cages with mosquito netting were constructed around each tray to prevent flying insects from accessing contaminated plants and soil.
Bacterial strains
Generic E. coli strains (TVS 353, TVS 354, and TVS 355) and Salmonella (Montevideo LJH-614, Michigan LJH-615, Saintpaul LJH-1262) were used in the inoculation cocktail. Generic E. coli strains had been previously isolated by Dr. Trevor Suslow's laboratory (University of California-Davis) from lettuce, irrigation water, and soil from the Salinas Valley and were adapted to be rifampicin-resistant by the Suslow laboratory. Salmonella strains were originally isolated from samples associated with foodborne outbreaks and had been previously adapted to be resistant to rifampicin by Dr. Linda Harris's laboratory (University of California-Davis).
Inoculum preparation
Stock cultures were stored at -80 ℃ in Tryptic Soy Broth (TSB; Neogen, Lansing, MI) with 40% glycerol. Frozen cultures of each strain were activated by transferring to TSB with incubation at 37 ℃ for 24 hours. For each strain, 0.1 ml of overnight culture was spread onto each of three Tryptic Soy Agar plates (TSA, Neogen) containing rifampicin (50 mg/L; Alfa Aesar, Ward Hill, MA; TSA + rif) and incubated at 37 ℃ for 22-26 hours. Bacterial lawns were harvested by adding 3 ml of 0.1% peptone water and scraping with a disposable cell spreader. Cell suspensions for each strain were collected separately and transferred to individual 15 ml sterile conical tubes. The cocktail was prepared by mixing 1 ml of each of the harvested lawns of the six strains into a 15-ml conical tube and held at 4 ℃ for up to 2 weeks. The stock cocktail solution was enumerated using standard serial dilution and spread plating techniques on Hektoen Enteric agar (Neogen) plates containing rifampicin (50 mg/L; HE + rif) to allow for differential enumeration (generic E. coli = yellow colonies; Salmonella = black colonies). The cocktail solution was diluted in 0.1% PW as necessary and 6 ml was then added to well water (19 L; private residential well, Philomath, OR) in a polypropylene carboy (U-Line, Pleasant Prairie, WI).
Soil inoculation
Inoculated well water (25 ml) was applied to each soil sample (400 g). To normalize inoculation levels, each sample was inoculated twice with two days between applications. Inoculated irrigation water samples (n = 3) were collected after each irrigation event and analyzed using standard serial dilution and spread plating techniques to verify contamination levels. Soils samples were maintained in the greenhouse for up to 86 days.
Microbial analysis
Soil samples were collected by aseptically pouring the complete soil content from a single cup into a sterile 710 ml Whirl-Pak filter bag (Nasco, Salida, CA). The 400-g sample was combined with 400 ml of 0.1% peptone water and mixed by hand for 20-30 seconds. Serial dilutions were prepared in 0.1% peptone with 0.1-1.0 ml being spread plated onto Hektoen Enteric Agar (Neogen, Lansing, MI) with rifampicin (50 mg/L; Alfa Aesar, Ward Hill, MA; HE + rif;). Plates were differentially enumerated for Salmonella and generic E. coli following incubation at 37 ℃ for 24 h. Five replicates were used to enumerate the bacterial density every 2 days in the first 20 days after the final contamination event and every 3 days thereafter.
When microbial counts from soil samples fell below the detection limit of 1 CFU/g for standard plating methods, a 96-well Most Probable Number (MPN) method was employed. The soil: peptone water mixture was aliquoted (1 ml) to 96 well deep well plates (VWR International, Radnor, PA). An additional 1ml of Lactose Broth (Neogen) containing rifampicin (50 mg/L; L + rif) was added to each well and incubated at 37 ℃ for 24 h. Following incubation, each well was spotted onto HE + rif plates using a 96 well tip comb. HE + rif plates were incubated 25 ℃ for 48 hours prior to evaluation. The lower incubation temperature was used to minimize colony overgrowth to improve differentiation of E. coli and Salmonella colonies. Five MPN samples were evaluated every three days. Qualitative positive results for generic E. coli and Salmonella were calculated as MPN/ml using Poisson distribution where d = -2.303/v log(s/n), n = total # of well, s = # of negative wells, and v = volume per well.
Once microbial counts fell below the detection limit of the MPN method (< 1 MPN/96 g) for two consecutive days, a qualitative total sample enrichment was employed. Ten soil samples (400 g) were collected weekly and enriched in 800 ml of L + rif with incubation at 37 ℃ for 24 h. Following incubation, enrichments were streaked onto HE + rif plates with subsequent incubated at 37 ℃ for 24 h. Generic E. coli presence was confirmed on Eosin Methylene Blue (EMB; Neogen) agar following incubation at 37 ℃ for 24 h. Soil analyses were discontinued when generic E. coli or Salmonella were no longer detected in any of the 10 enrichment samples (up to 86 days).
Results and Discussion
Surface waters in Oregon that are used for irrigation of produce vary significantly in their microbial quality. For the purposes of this study, we were interested in evaluating the persistence of generic E. coli in soils that had been irrigated with water of exceptionally poor microbial quality. There are areas within the state of Oregon that utilize surface water for irrigating root crops with historic E. coli levels that occasionally exceed 10,000 MPN/100 ml (3 log CFU/ml) (personal communication). This contamination level was the target for our "low inoculation" water. Generic E. coli and Salmonella concentrations in the low inoculation irrigation water averaged 2.70 ± 0.09 log CFU/ml and 2.76 ± 0.12 log CFU/ml, respectively. To evaluate the survival of microbial contamination in soils under a "worst-case" scenario, our "high inoculation" water was targeted at a 100-fold increase of the "low" level. Generic E. coli and Salmonella concentrations in the high inoculation irrigation water were 4.59 ± 0.06 log CFU/ml and 4.93 ± 0.04 log CFU/ml, respectively. Two applications of contaminated water (2 × 25 ml) to soil (400 g) resulted in generic E. coli and Salmonella levels in high inoculum soil samples of 3.36 ± 0.31 and 3.60 ± 0.21 log CFU/g, respectively. Generic E. coli and Salmonella levels in all low inoculum soil samples on day 0 were 2.09 ± 0.45 and 2.39 ± 0.45 log CFU/g, respectively.
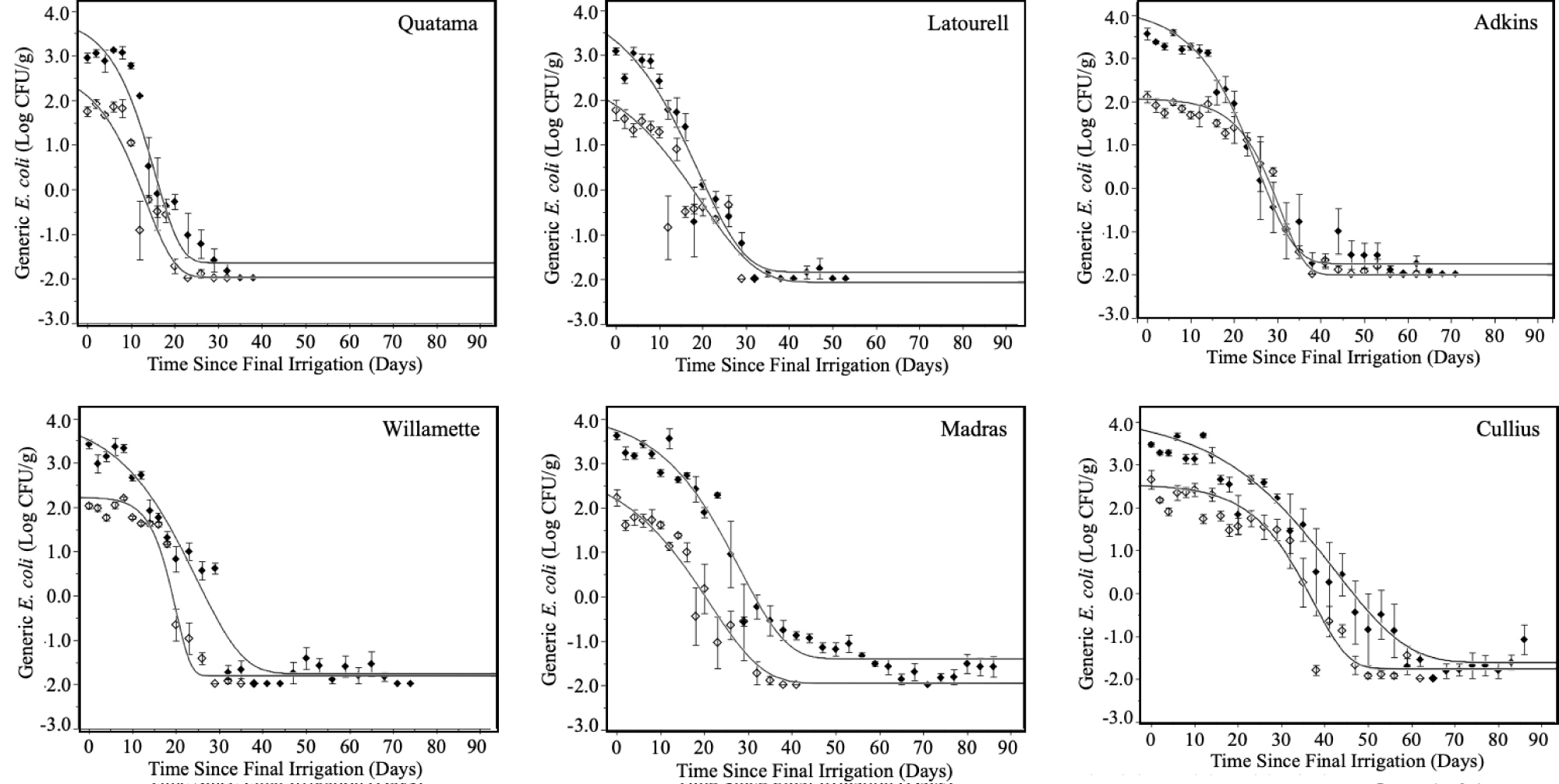
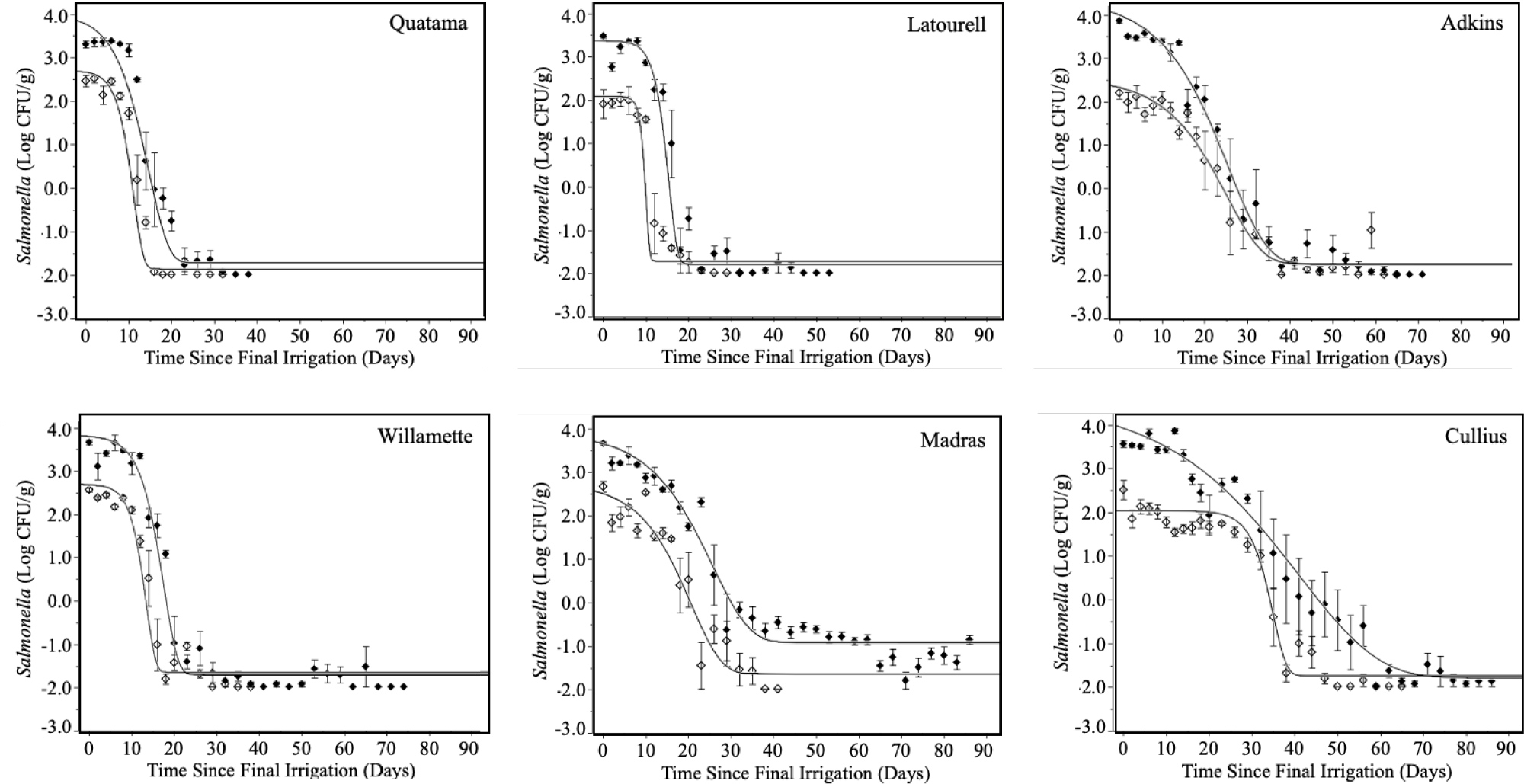
Survival and persistence of high inoculum and low inoculum generic E. coli and Salmonella in different soils are shown in (Figure 1 and Figure 2). All soil types revealed similar initial contamination levels at day 0. Generic E. coli and Salmonella survival patterns were comparable in all soil types at both inoculum levels. This was in good agreement with previous research showing similar survival curves for generic E. coliand Salmonella Typhimurium in diverse agricultural soils [13,21,22]. Both E. coli and Salmonella populations were stable for a period of days, followed by a period of sharp decline, with a low level of survivors persisting for extended periods of time (≥ 80 days). The same pattern of bacterial survival has been demonstrated previously in a variety of low moisture environments, including agricultural soils [8,10,11,23-26]. Soils inoculated at the higher contamination level harbored low numbers of surviving cells for longer periods of time than soils inoculated at the lower inoculation level. With the exception of Quatama, survivors of E. coli and Salmonella were detected in at least 20% of soil samples after > 80 days of irrigation with highly contaminated water (Table 2). These results are in agreement with previous reports [8,11].
Individual soils revealed significant differences in their ability to support survival of the inocula. Quatama revealed the most inhibitory effect on generic E. coli and Salmonella with rapid die-off occurring in the first 20 days (Figure 1 and Figure 2) and no survivors being detected at 49 and 69 days at low and high inoculum levels, respectively (Table 2). Generic E. coli and Salmonella persisted only slightly better in Latourell and Willamette soils as compared to Quatama. The enhanced persistence in Latourell and Willamette soils was most clearly demonstrated in soil samples inoculated at high levels where 20-50% of soil samples remained positive for at least one of the inoculated species through the end of the study (84 days post-inoculation; Table 2). The rates of reduction of the inocula in Madras and Adkins soils were significantly lower when compared with those of Quatama, Latourell, and Willamette, taking nearly 40 days to complete the rapid die-off period. Cullius was the most supportive of the survival and persistence of generic E. coli and Salmonella, taking nearly 60 days to complete the die-off period for highly contaminated soils (Figure 1and Figure 2). The correlations of specific soil characteristics with the survival and persistence of E. coli have been reported previously [9,27-31]. Different factors have been reported to have significant impact on pathogen survival including soil type, including soil texture (clay, silt, sand), soil pH, organic matter, exchangeable bases, nitrogen, temperature (static and variable), solar radiation, soil moisture/water activity, matric potential, oxidation-reduction potential, clay mineral type, microbial biomass carbon (MBC), microbial interactions (predation/competition; relative abundance of Acidobacteria and Chloroflexi), contamination level, and method of contamination [10,15,27,32-34].
The data of physical and chemical properties of agricultural soils from diverse growing regions of Oregon (Table 1) revealed that there are substantial differences in physical and chemical characteristics of these soils such as water availability, organic matter contents, cation exchange capacity, mineral contents, pH, and soil texture (clay%, silty% and sand%). The differences in these characteristics may explain the differences in survival and persistence of generic E. coli and Salmonella among different tested soils. These data (Table 1) indicate no difference in available water between Quatama (the most inhibiting soil) and Cullius (the most supportive soil), therefore, the water content does not play a significant factor that affect the persistence of the pathogens. This was in disagreement with numerous researchers who suggested soil moisture as a factor that influences the survival of E. coli and other bacteria in soil systems [21,27,32,35-37]. The moisture in soil systems is a dynamic, dependent variable and it is greatly influenced by experimental design and environmental conditions; therefore, it cannot be considered as a main factor that affect bacterial persistence in the soil. Moreover, studies provided information related to soil moisture content used a wide variety of terms (percent moisture, percent water-holding capacity, soil moisture tension, degree of saturation) and often leave out critical descriptive elements to allow for the reader to compare between studies [27]. Several authors have attempted to evaluate bacterial survival in soil systems using designs that maintain moisture (closed/sealed systems); however, results from these systems were the least practical for translating results to agricultural settings. Others have controlled the moisture by adjusting the initial moisture level of the soil to achieve a relative field capacity across all soils and they revealed differing conclusions about the impact of initial moisture on bacterial survival [8,32]. In the current study, the chosen approach was similar to that of Fremaux, et al. [15] and applied the same water: soil ratio across all soil types to evaluate the bacterial survival as a function of the soils' innate characteristics. Throughout the course of the current study, soil samples dried as a function of temperature and humidity of the greenhouse environment. The study was not designed to correct for starting moisture of the soil samples; nor was it designed to focus on the moisture loss rates for different soil types. Regardless, moisture loss over time was apparent and differences in the rates of moisture loss between soils likely contributed to bacterial inactivation rates, particularly during periods of exponential reduction.
Soil texture has been recognized as an important factor that can influence the survival of pathogens in soils. The data in (Table 1) revealed that Quatama (the most inhibiting soil) contains a higher percentage of sand; however, Cullius (the most supportive soil) contains a higher percentage of clay. Sandy soils contain a predominance of non-cohesive particles which leads to a poor retention of nutrients and a lower water holding capacity. These characteristics have been attributed to shorter periods of survival for enteric pathogens in sandy soils [10,12]. However, other studies have demonstrated increased survival and long-term persistence in sandier soils suggesting that moisture status plays an insignificant role when evaluating the potential for long-term persistence of E. coli in dry soil systems [11,15,16,33]. However, clay content of soils has been correlated with increased survival of enteric pathogens in several studies [12,15,29,38-41]. Clay is the smallest particle size fraction (< 0.002 mm) providing a large amount of surface area; therefore, soils high in clay may favor the adsorption of bacteria to particles and create a physical barrier (pore size) against microbial predators and parasites [39]. Supplementation of a sandy brown soil with three clay minerals (montomorillonite/smectite, kaolinite, and illite) was shown to improve the survival of Listeria monocytogenes, E. coli O157:H7, and Salmonella Dublin [40]. Brennan, et al. [40] observed that each clay mineral had a unique impact on the cation exchange capacity and surface area of the soil and the survival of pathogens was improved with the addition of all clay minerals; however, montmorillonite revealed the greatest increase and kaolinite revealed the least improvement in survival of pathogens. These studies demonstrate the impact of distinct clay minerals on bacterial survival in soils; however, soils contain a mixture of clay minerals that are often not well defined. NRCS classifies soils based on predominant clay mineral component using x-ray diffraction [42]; however, many soils are classified as "mixed" which provides little opportunity to evaluate the relative clay mineral composition of these soils. Of the soils investigated in this study, Cullius had the highest composition of clay (39.1%) and was classified as smectitic mineralogy. All remaining soils were < 25% clay and classified as having mixed mineralogy. These classifications of Cullius support the significantly improved survival of generic E. coli and Salmonella compared to the other Oregon soils investigated in this study.
One of the most important factors that likely influences the survival and persistence of pathogens is the pH of the soil. Quatama has the lowest pH value (5.1, acidic), which may explain the most inhibitory effect of this soil type, whereas Cullius has the highest pH value (6.6, nearly neutral), which explain the most supportive effect of this soil type. It has been observed previously that E. coli survives longer in soils at neutral to slightly alkaline pH (6.0-8.5) compared to acidic soils [9,26,27,37,43,44]. The survival of E. coli O157:H7 was evaluated in 14 soil types from eastern China and the survival of the pathogen was superior in slightly alkaline soils (pH 8.4-8.6) and poor in highly acidic soils (pH 4.2-4.7) [26]. The pH of environmental systems has an inherent impact on the stress of bacterial contaminants and affect the bacterial adsorption characteristics of clay minerals and other associated soil chemistry [36,41,45].
Other components of the soil, such as organic matter and mineral contents, which may affect the survival of the pathogens are presented in (Table 1). Although Quatama has higher organic matter contents in comparison to other soil types, it revealed the most inhibitory effect. However, Cullius has the lowest organic matter contents and it revealed the most supportive effect to the pathogens. Quatama contains a higher amount of NO3-N and lower mineral content, meanwhile, Cullius has lower NO3-N and higher mineral content. These components may have significant impacts on the electrochemistry of the soil system which will influence the moisture retention of the soil, adsorption of bacteria to the soil, and proximity of bacteria to chemical stresses (production of reactive oxygen species) [39,41]. Application of biological soil amendments provides an additional source for contamination; however, the addition of "safe" biological soil amendments will likely increase the survival of enteric pathogens in agricultural soils. Persistence of enteric bacteria has been demonstrated in soil systems with low levels of E. coli O157:H7 and E. coli O26 being detected beyond 270 days post-inoculation in various manure-amended soils [15,33].
Conclusion
From the current study, it can be concluded that the survival and persistence of generic E. coli and Salmonella were affected by the soil type. Quatama revealed the most inhibitory effect; however, Cullius revealed the most supportive effect on the survival and persistence of these microorganisms. The physical and chemical properties of soils such as water availability, soil texture, cation exchange capacity, pH, organic matter and mineral contents are important factors which may affect the survival and persistence of microorganisms in different soils. This study demonstrated the long-term risk posed by the survival of foodborne pathogens introduced to soil by contaminated irrigation water. The results of this study may help produce growers in identifying mitigation strategies for reducing pathogens in soil and preventing contamination of edible crops that may be in direct contact with soil, particularly those soils that are likely to support survival and persistence of enteric pathogens.
Acknowledgments
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors. The authors would like to thank Marc Anderson and Michael Bondi (North Willamette Research & Extension Center (NWREC), Aurora, OR), Hoyt Downing and Carol Tollefson (Central Oregon Agricultural Research Center (COARC), Madras, OR), and Tim Weinke (Hermiston Agricultural Research & Extension Center, Hermiston, OR) for supplying soil used in this study. Thank you to Dr. Linda Harris at University of California, Davis for supplying the strains used in this study. We would also like to thank Jim Ervin and Gloria O'Brien of OSU's Greenhouse Team for their guidance within the greenhouse. Thank you to Dr. Mark Daeschel for providing the well water for this study. Chris Letchworth, Daniel Wright, Joey Minarsich, and Amy Emch deserve special thanks for their efforts to support of this research project.
References
- Gould LH, Walsh K, Vieira AR, et al. (2013) Surveillance for foodborne disease outbreaks - United States, 1998-2008. MMWR Surveill Summ 62: 1-34.
- Painter JA, Hoekstra RM, Ayers T, et al. (2013) Attribution of foodborne illnesses, hospitalizations, and deaths to food commodities by using outbreak data, United States, 1998-2008. Emerg Infect Dis 19: 407-415.
- Chen Parker C, McKenna C, Wise M, et al. (2017) Geospatial mapping of early cases in multistate foodborne disease outbreaks: A strategy to expedite identification of contaminated imported produce, United States, 2006 to 2013. J Food Prot 80: 1821-1831.
- United States Food and Drug Administration (2015a) Standards for the growing, harvesting, packing, and holding of produce for human consumption. Final Rule. Fed Regist 80: 74353-74672.
- United States Food and Drug Administration (2015b) Current good manufacturing practice, hazard analysis, and risk-based preventive controls for human food. Fed Regist 80: 55908-56168.
- United States Food and Drug Administration (2019) Standards for the growing, harvesting, packing, and holding of produce for human consumption; extension of compliance dates for subpart E. Final rule. Fed Regist 84: 9706-9714.
- Isaacs S, Aramini J, Ciebin B, et al. (2005) An international outbreak of salmonellosis associated with raw almonds contaminated with a rare phage type of Salmonella Enteritidis. J Food Prot 68: 191-198.
- Danyluk MD, Nozawa-Inoue M, Hristova KR, et al. (2008) Survival and growth of Salmonella Enteritidis PT 30 in almond orchard soils. J Appl Microbiol 104: 1391-1399.
- Sjogren RE (1994) Prolonged survival of an environmental Escherichia coli in laboratory soil microcosms. Water, Air Soil Pollut 75: 389-403.
- Fenlon DR, Ogden ID, Vinten A, et al. (2000) The fate of Escherichia coli and E. coli O157 in cattle slurry after application to land. Symp Ser Soc Appl Microbiol 88: 149S-156S.
- Cools D, Merckx R, Vlassak K, et al. (2001) Survival of E. coli and Enterococcus spp. derived from pig slurry in soils of different texture. Appl Soil Ecol 17: 53-62.
- Lau MM, Ingham SC (2001) Survival of faecal indicator bacteria in bovine manure incorporated into soil. Lett Appl Microbiol 33: 131-136.
- Natvig EE, Ingham SC, Ingham BH, et al. (2002) Salmonella enterica serovar Typhimurium and Escherichia coli contamination of root and leaf vegetables grown in soils with incorporated bovine manure. Appl Environ Microbiol 68: 2737-2744.
- Ibekwe AM, Watt PM, Shouse PJ, et al. (2004) Fate of Escherichia coli O157: H7 in irrigation water on soils and plants as validated by culture method and real-time PCR. Can J Microbiol 50: 1007-1014.
- Fremaux B, Prigent Combaret C, Delignette Muller ML, et al. (2008) Persistence of Shiga toxin-producing Escherichia coli O26 in various manure-amended soil types. J Appl Microbiol 104: 296-304.
- Bolton DJ, Monaghan A, Byrne B, et al. (2011) Incidence and survival of non-O157 verocytotoxigenic Escherichia coli in soil. J Appl Microbiol 111: 484-490.
- United states department of agriculture - National resources conservation service (USDA-NRCS) (2019) Prime farmland list for Oregon.
- Emch AW, Waite-Cusic JG (2016) Conventional curing practices reduce generic Escherichia coli and Salmonella spp. on dry bulb onions produced with contaminated irrigation water. Food Microbiol 53: 41-47.
- Huddleston J (1979) Special Report 535: Soils of oregon: Their classification, taxonomic relationships, and physiography.
- United States Department of Agriculture - National Resources Conservation Service (1999) Soil Taxonomy: A basic system of soil classification for making and interpreting soil surveys, Second Edition.
- Chandler DS, Craven JA (1980) Relationship of soil moisture to survival of Escherichia coli and Salmonella Typhimurium in soils. Aust J Agric Res 31: 547-555.
- Plachá I, Venglovský J, Sasáková N, et al. (2001) The effect of summer and winter seasons on the survival of Salmonella Typhimurium and indicator micro-organisms during the storage of solid fraction of pig slurry. J Appl Microbiol 91: 1036-1043.
- Giles N, Hopper SA, Wray C (1989) Persistence of S. Typhimurium in a large dairy herd. Epidemiol Infect 103: 235-241.
- Henschke RB, Henschke EJ, Schmidt FRJ (1991) Monitoring survival and gene transfer in soil microcosms of recombinant Escherichia coli designed to represent an industrial production strain. Appl Microbiol Biotechnol 35: 247-252.
- Islam M, Morgan J, Doyle MP, et al. (2004b) Fate of Escherichia coli O157: H7 in manure compost-amended soil and on carrots and onions grown in an environmentall controlled growth chamber. J Food Prot 67: 574-578.
- Wang H, Ibekwe AM, Ma J, et al. (2014) A glimpse of Escherichia coli O157: H7 survival in soils from eastern China. Sci Total Environ 476-477: 49-56.
- Reddy KR, Khaleel R, Overcash MR (1981) Behavior and transport of microbial pathogens and indicator organisms in soils treated with organic wastes. J Environ Qual 10: 255-266.
- Vandepitte V, Quataert P, de Rore H, et al. (1995) Evaluation of the Gompertz function to model survival of bacteria introduced into soils. Soil Biol Biochem 27: 365-372.
- Franz E, Semenov A V, Termorshuizen AJ, et al. (2008) Manure-amended soil characteristics affecting the survival of E. coli O157: H7 in 36 dutch soils. Environ Microbiol 10: 313-327.
- Yao Z, Wei G, Wang H, et al. (2013) Survival of Escherichia coli O157: H7 in soils from vegetable fields with different cultivation patterns. Appl Environ Microbiol 79: 1755-1756.
- Wang H, Ryser ET (2014) Salmonella transfer during pilot plant scale washing and roller conveying of tomatoes. J Food Prot 77: 380-387.
- Mubiru DN, Coyne MS, Grove JH (2000) Mortality of Escherichia coli O157: H7 in two soils with different physical and chemical properties. J Environ Qual 29: 1821-1825.
- Jiang X, Morgan J, Doyle MP (2002) Fate of Escherichia coli O157: H7 in manure-amended soil. Appl Environ Microbiol 68: 2605-2609.
- Jacobsen CS, Bech TB (2012) Soil survival of Salmonella and transfer to freshwater and fresh produce. Food Res Int 45: 557-566.
- Kibbey HJ, Hagedorn C, Mccoy EL (1978) Use of fecal streptococci as indicators of pollution in soil. Appl Environ Microbiol 35: 711-717.
- Tate III RL (1978) Cultural and environmental factors affecting the longevity of Escherichia coli in histosols. Appl Enviromental Microbiol 35: 925-929.
- Jamieson RC, Gordon RJ, Sharples KE, et al. (2002) Movement and persistence of fecal bacteria in agricultural soils and subsurface drainage water: A review. Can Biosyst Eng 44: 1.1-1.9.
- Howell JM, Coyne MS, Cornelius PL (1996) Effect of sediment particle size and temperature on fecal bacteria mortality rates and the fecal coliform/fecal streptococci ratio. J Environ Qual 25: 1216-1220.
- Santamaría J, Toranzos GA (2003) Enteric pathogens and soil: A short review. Int Microbiol 6: 5-9.
- Brennan FP, Moynihan E, Griffiths BS, et al. (2014) Clay mineral type effect on bacterial enteropathogen survival in soil. Sci Total Environ 468-469: 302-305.
- Liu X, Gao C, Ji D, et al. (2017) Survival of Escherichia coli O157: H7 in various soil particles: Importance of the attached bacterial phenotype. Biol Fertil Soils 53: 209-219.
- U.S. Natural Resources Conservation Service Soil Survey Staff (2014) Kellogg Soil Survey Laboratory Methods Manual.
- Cuthbert W, Panes JJ, Hill EC (1955) Survival of Bacterium coli type i and Streptococcus faecalis in soil. J Appl Microbiol 18: 408-414.
- Ellis JR, McCalla TM (1978) Fate of pathogens in soils receiving animal wastes - a review. Trans ASAE 309-313.
- Ma J, Ibekwe AM, Yi X, et al. (2011) Persistence of Escherichia coli O157: H7 and its mutants in soils. PLoS One 6: e23191.
Corresponding Author
Joy Waite-Cusic, Department of Food Science and Technology, 100 Wiegand Hall, Oregon State University, Corvallis, Oregon 97331, USA, Tel: +1(541)737-6825
Copyright
© 2020 Emch AW, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.